-Search query
-Search result
Showing 1 - 50 of 121 items for (author: shek & j)

EMDB-18373: 
cryo-EM structure of apo Clostridioides difficile toxin B
Method: single particle / : Kinsolving J, Bous J, Structural Genomics Consortium (SGC)

EMDB-18374: 
cryo-EM structure complex of Frizzled-7 and Clostridioides difficile toxin B
Method: single particle / : Kinsolving J, Bous J

PDB-8qen: 
cryo-EM structure of apo Clostridioides difficile toxin B
Method: single particle / : Kinsolving J, Bous J, Structural Genomics Consortium (SGC)

PDB-8qeo: 
cryo-EM structure complex of Frizzled-7 and Clostridioides difficile toxin B
Method: single particle / : Kinsolving J, Bous J

EMDB-35304: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3A) complex
Method: single particle / : Xie T, Gong X

EMDB-35306: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3A-N71A) complex
Method: single particle / : Xie T, Gong X

EMDB-35310: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3D) complex
Method: single particle / : Xie T, Gong X

PDB-8iaj: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3A) complex
Method: single particle / : Xie T, Gong X

PDB-8iak: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3A-N71A) complex
Method: single particle / : Xie T, Gong X

PDB-8iam: 
Cryo-EM structure of the yeast SPT-ORM2 (ORM2-S3D) complex
Method: single particle / : Xie T, Gong X

EMDB-28965: 
Glutamine synthetase from Pseudomonas aeruginosa, filament double-unit in compressed conformation
Method: single particle / : Phan IQ, Staker B, Shek R, Moser TH, Evans JE, van Voorhis WC, Myler PJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8fbp: 
Glutamine synthetase from Pseudomonas aeruginosa, filament double-unit in compressed conformation
Method: single particle / : Phan IQ, Staker B, Shek R, Moser TH, Evans JE, van Voorhis WC, Myler PJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-16569: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003
Method: single particle / : Farrell B, Higgins MK

EMDB-16570: 
PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003
Method: single particle / : Farrell B, Higgins MK

EMDB-16635: 
PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 - maps for local refinement on PfRIPR
Method: single particle / : Farrell B, Higgins MK

EMDB-16636: 
PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 consensus maps
Method: single particle / : Farrell B, Higgins MK

EMDB-16637: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 - consensus map
Method: single particle / : Farrell B, Higgins MK

EMDB-16638: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 - local refinement on PfRH5
Method: single particle / : Farrell B, Higgins MK

EMDB-16639: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 - local refinement on PfRIPR
Method: single particle / : Farrell B, Higgins MK

EMDB-16640: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003 - map with additional PfRIPR tail density
Method: single particle / : Farrell B, Higgins MK

PDB-8cdd: 
PfRH5-PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003
Method: single particle / : Farrell B, Higgins MK

PDB-8cde: 
PfCyRPA-PfRIPR complex from Plasmodium falciparum bound to antibody Cy.003
Method: single particle / : Farrell B, Higgins MK

EMDB-40671: 
Cryo-EM structure of Enoyl-CoA hydratase from Mycobacterium smegmatis
Method: single particle / : Shek R, Quispe J, Staker B, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8soy: 
Cryo-EM structure of Enoyl-CoA hydratase from Mycobacterium smegmatis
Method: single particle / : Shek R, Quispe J, Staker B, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-33873: 
Cryo-EM structure of the dimeric atSPT-ORM1 complex
Method: single particle / : Xie T, Liu P, Gong X

EMDB-33874: 
Cryo-EM structure of the monomeric atSPT-ORM1 complex
Method: single particle / : Xie T, Liu P, Gong X

EMDB-33875: 
Cryo-EM structure of the monomeric atSPT-ORM1 (ORM1-N17A) complex
Method: single particle / : Xie T, Liu P, Gong X

EMDB-33876: 
Cryo-EM structure of the monomeric atSPT-ORM1 (LCB2a-deltaN5) complex
Method: single particle / : Xie T, Liu P, Gong X

PDB-7yjk: 
Cryo-EM structure of the dimeric atSPT-ORM1 complex
Method: single particle / : Xie T, Liu P, Gong X

PDB-7yjm: 
Cryo-EM structure of the monomeric atSPT-ORM1 complex
Method: single particle / : Xie T, Liu P, Gong X

PDB-7yjn: 
Cryo-EM structure of the monomeric atSPT-ORM1 (ORM1-N17A) complex
Method: single particle / : Xie T, Liu P, Gong X

PDB-7yjo: 
Cryo-EM structure of the monomeric atSPT-ORM1 (LCB2a-deltaN5) complex
Method: single particle / : Xie T, Liu P, Gong X
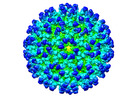
EMDB-26945: 
Cryo-EM structure of SINV/EEEV in complex with Fab fragment of a moderately/weakly neutralizing human antibody IgG-21
Method: single particle / : Bandyopadhyay A, Klose T, Kuhn RJ
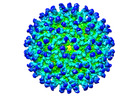
EMDB-26946: 
Cryo-EM structure of SINV/EEEV in complex with Fab fragment of a moderately/weakly neutralizing human antibody IgG-94
Method: single particle / : Bandyopadhyay A, Klose T, Kuhn RJ

EMDB-26947: 
Cryo-EM structure of SINV/EEEV in complex with Fab fragment of a potently neutralizing human antibody IgG-106
Method: single particle / : Bandyopadhyay A, Klose T, Kuhn RJ

PDB-7v0n: 
Cryo-EM structure of SINV/EEEV in complex with Fab fragment of a moderately/weakly neutralizing human antibody IgG-21
Method: single particle / : Bandyopadhyay A, Klose T, Kuhn RJ

PDB-7v0o: 
Cryo-EM structure of SINV/EEEV in complex with Fab fragment of a moderately/weakly neutralizing human antibody IgG-94
Method: single particle / : Bandyopadhyay A, Klose T, Kuhn RJ

PDB-7v0p: 
Cryo-EM structure of SINV/EEEV in complex with Fab fragment of a potently neutralizing human antibody IgG-106
Method: single particle / : Bandyopadhyay A, Klose T, Kuhn RJ

EMDB-28092: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-093
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28090: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-040
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28091: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-045
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28093: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-156
Method: single particle / : Shek J, Callaway H, Li H, Yu X, Saphire EO

EMDB-28094: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-234
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28095: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-260
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28096: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-279
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28097: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-290
Method: single particle / : Yu X, Callaway H, Li H, Shek J, Saphire EO
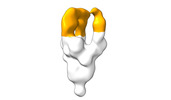
EMDB-28098: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-294
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28099: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-295
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28100: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-299
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28102: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-334
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
